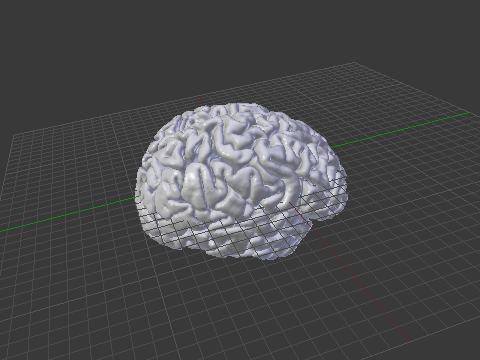
Computer graphics (CG) applications, such as Blender, Maya or 3D Studio, doesn’t come with filters that allow to directly import or export FreeSurfer meshes. To import brain surfaces to these applications, converting first to an intermediate file format solves the problem. Below is an example of a brain imported into Blender.
The mris_convert, from FreeSurfer, allows conversion to stereolithography format (STL). However, not all CG applications read natively from STL files. Moreover, other formats, such as Wavefront OBJ and Stanford PLY have simpler structure — yet allowing features that will be discussed here in forthcoming articles.
Conversion from FreeSurfer to Wavefront OBJ
A simple procedure to convert a FreeSurfer surface to Wavefront OBJ (ASCII) is:
1) Use mris_convert to convert from FreeSurfer binary to ASCII:
mris_convert lh.pial lh.pial.asc
2) A suggestion is to rename the .asc file to .srf, to differentiate it from other files that also have .asc extension and have a different internal structure.
mv lh.pial.asc lh.pial.srf
3) Use the script srf2obj to convert from .srf to .asc:
srf2obj lh.pial.srf > lh.pial.obj
The symbol “>” is necessary so that the output from srf2obj is redirected to a new file.
Conversion from Wavefront OBJ to FreeSurfer
To convert back to FreeSurfer, a similar strategy is used.
1) Use the script obj2srf to generate new .srf files (here directly named as .asc, as expected by mris_convert)
obj2srf lh.pial.obj > lh.pial.asc
2) Use mris_convert:
mris_convert lh.pial.asc lh.pial
Download
Both scripts are available here: srf2obj and obj2srf. Right-click and save, then make them executable.
Requirements
The scripts use gawk as the interpreter, and expects to find it on its usual location (/bin/gawk). If in your system gawk is installed elsewhere, modify the first line accordingly.
If you don’t have gawk, you can get it from the repository of your Linux distribution. For Mac, gawk is available in MacPorts and Fink. Alternatively, you may replace gawk with awk itself, nawk, or any other awk implementation that works.

Pingback: Braindering with ASCII files | Brainder.
Pingback: Importing FreeSurfer subcortical structures into Blender | Brainder.
Hello! This is a great site, by the way. I’ve been working on visualization of neuroscientific visualization with blender for a little while now, but only using .stl files. Exciting to meet someone else working on this!
Anyway, I’ve been trying to get your srf2obj script working, but it bombs:
bash: /mnt/home/p4u1/Shellscripts/BASH_Scripts/srf2obj.sh: line 3: syntax error near unexpected token `(‘
bash: /mnt/home/p4u1/Shellscripts/BASH_Scripts/srf2obj.sh: line 3: `BEGIN { if (ARGV[1] == “”) { ‘
My gawk is in /usr/bin, and I changed the first line accordingly, but admit I am not too familiar with (g)awk (mainly just use it for separating fields. If I comment out the usage, I get:
}: command not found
NR: command not found
NR: command not found
I assume I’m just not invoking the correct interpreter, but any help would be appreciated.
Thanks,
Paul
Dear Paul,
Yes, apparently the interpreter isn’t right. It should work out of the box with gawk, and it should also work with other awk implementations (such as nawk). Double check that you have really gawk (not simply a symlink pointing to some other implementation) and that the first line of the script is correct. If you download the file from Windows, make sure you use the right-click, then “save as”, otherwise Windows may introduce some undesired characters.
All the best,
Anderson
Ah-HA! Now I have it: it looks like your script requires the ?h.pial.srf file to be in the same directory as the script — I had been giving it the full path.
Works like a charm :)
Thanks,
Paul
Hi Paul,
In fact, this shouldn’t be an issue. I can run here with the script located, say, in directory A, with the files in some other directory B, and calling all from yet another directory C. In any case, glad it worked, and thanks for the feedback.
All the best,
Anderson
Pingback: Splitting the cortical surface into independent regions | Brainder.
Hey there, I’m having a little trouble creating subcortical surfaces. I’ve created the freesurfer meshes using aseg2srf but using the above commands to convert to a CG software compatible file is doing weird things. mris_convert creates a stream of “cannot allocate memory” type messages in terminal window and essentially grinds things to a halt. Using srf2obj the output file is a tgif file 0 bytes in size… any ideas? Maybe aseg2srf didn’t work properly?
Hi Will,
This looks strange. If the final outputs from aseg2srf are files with extension .srf, which when opened in a simple text editor look like an orderly set of vertex coordinates, then it should have worked. Errors related to memory allocation seem something with your particular system. Send me an email via the “About” form and we can try to sort it out.
All the best,
Anderson
Hi,
I’m having the same issue where srf2obj creates a lh.pial.obj that is Zero bytes in size. Did you ever find a solution?
Thanks!
Hi,
Did you follow all the steps? Make sure the input file is converted to ASCII before running srf2obj.
All the best,
Anderson
I’m using a Mac and I followed your steps. Maybe my mistake is in making the script executable? I used “chmod +x” to do so. Should the script be a .text or .sh? I’m not very well versed in programming.
I ran the script through awk. I have Fink installed on my computer. The problem persists though I have tried any things.
On Thursday, October 30, 2014, Brainder. wrote:
> A. M. Winkler commented: “Hi, Did you follow all the steps? Make sure > the input file is converted to ASCII before running srf2obj. All the best, > Anderson”
Hi,
Yes, the script should be executable. It can have any extension, or no extension at all, as in the example in the page. Using awk or gawk should be equivalent, although some systems (including the newer Macs) may use older, BSD-licensed, implementations of awk.
Did you remember to use the > sign? (see the example).
If it still fails, put the file somewhere online where I can download and I’ll have a look.
Cheers,
Anderson
Sorry about the late reply. I did have this problem a while ago but have set up a new server since then and have not replicated it. I’m an amateur myself, Anderson do you think permissions or ownership of directories or the script could cause the problem? This has been my first point of call…
Yes, the script should be executable, no doubt. If not, it’s going to fail.
Pingback: Manual to print your own brain |
I have a question, already apologizing if it is obvious and I’m just not able to see it. I was trying to create an .obj from the inflated hemispheres. That works fine for each seperate hemisphere. However, trying to combine them, both hemispheres end up overlaying each other, looking like both their origins are set to 0 so they overlap. (Using the merge surfs option from mris_convert did not help)
How do you manage to comine the lh.obj and the rh.obj surface to a whole brain surface mesh?
Thanks a lot for your help.
Hi Lisa,
Each hemisphere stays in a separate file, and when the inflation process happens, there is no constraint on intersection between left and right.
To deal with this issue you’d have to manually apply some translation along the x-axis. There are many ways to do this: one is opening the surface files in, e.g., Matlab, adding or subtracting some constant value to the x-coordinate on each hemisphere (positive or negative, depending on the side), then saving.
Hope this helps!
All the best,
Anderson