Suppose you run a statistical test for each of the regions of the parcellated brain produced by FreeSurfer. It can be either surface area comparison between groups, or maybe differences in thickness. For each region, you may obtain a statistical score and likely a p-value as well. How to display these results in a color-coded brain model, still inside FreeSurfer?
The process is consists of three steps:
- Create of a table with your results, in a format that FreeSurfer can read.
- Embed the table in an annotation file.
- Display in
tksurfer.
1. Create a table
The table must contain not the actual scalars, but instead rgb triplets. This can be done using a simple Octave or matlab script. Since often these results are tabulated as spreadsheets, an alternative, straightforward way is using software like LibreOffice, which gives instantaneous graphical feedback of how the resulting table looks like.
An example of such a spreadsheet is this: colormap.ods. The data go into the sheets named lh.data and rh.data. For structures that are not supposed to display any color, put an off-range value below the minimum value of the other regions (e.g. -1 if the scalars are all positive). Do not add or remove structures.
The sheet named colormap contains the key between the scalar values and the actual rgb colors. In this sheet, the first column (column A) is simply a rank number, used in the calculations; the second column (col. B) contains the range of values between 0 and max, regularly spaced. The remaining three columns (C, D and E) contains the rgb triplets to be assigned to each value in column B. For each scalar to be shown, the closest number has its corresponding rgb used.
The sheets named lh.aparc.annot.ctab and rh.aparc.annot.ctab contain the resulting tables that will be used for actual display purposes. Each contain an index number, starting from 0, the structure name as used in FreeSurfer, the rgb colors, and a fourth value that is called “flag” and is usually zero.
In most cases, only the numerical values in lh.data and rh.data have to be changed. In some occasions, it may be necessary to change the range to be shown (i.e., column B on the colormap sheet), or the colormap itself (columns C, D and E). Different colormaps can be generated, for instance, using Octave. In the file used here, the colormap is the “spectrum”, also known as “jet”, shown below:
Finally, FreeSurfer cannot read binary spreadsheets like this. The file has to be converted to simple values separated by space. In LibreOffice or OpenOffice, there is a very simple way of doing this. Go to File -> Save As… In the File type, choose Text CSV, and make sure to mark the option Edit Filter Settings:
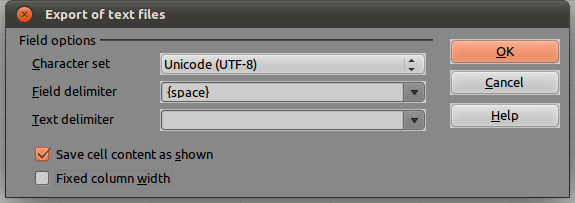
On the next window, as Field Delimiter, choose {space}. As Text delimiter, leave blank, as shown below:
The resulting file, saved to the disk, should look like this (click here for the full file):
0 bankssts 92 255 163 0
1 caudalanteriorcingulate 143 255 112 0
2 caudalmiddlefrontal 0 92 255 0
[...]
34 transversetemporal 255 133 0 0
35 unknown 160 160 160 0
2. Embed the table into annotation file
Once a table has been produced, it can be embedded into the annotation file that defines the the vertices that belong to each region. The annotation files produced by FreeSurfer already contain a default color table, which can be replaced. This can be accomplished by the Octave function replace_ctab.m. This function needs Octave 3.4.0+ or matlab. Type help replace_ctab at the prompt to get usage information. Here is an example:
replace_ctab('${SUBJECTS_DIR}/bert/label/lh.aparc.annot',...
'${SUBJECTS_DIR}/bert/label/lh.aparc.annot.ctab.csv',...
'${SUBJECTS_DIR}/bert/label/lh.aparc.annot.new');
In this example, the original annotation is stored in the file ${SUBJECTS_DIR}/bert/label/lh.aparc.annot, the new color table, as saved in LibreOffice, is in the file ${SUBJECTS_DIR}/bert/label/lh.aparc.annot.ctab. The new annotation file, with the new colortable, will be in the file ${SUBJECTS_DIR}/bert/label/lh.aparc.annot.new. Notice that for the function replace_ctab to work, the directory ${FREESURFER_HOME}/matlab must be in the Octave or matlab path.
3. Display with tksurfer
Once the new annotation file has been produced, display the result is straightforward:
tksurfer bert lh pial -annotation lh.aparc.annot.new
The result should be like the brain shown at the beginning of this article.
Update (01.Aug.2013): In more recent FS versions, the “lh” in front of the annotation file can be omitted, as it’s already specified in the same command line (thanks to Krishna Pancholi, Olin Neuropsychiatric Research Center, for the tip.). This affects only the last command line, which becomes then:
tksurfer bert lh pial -annotation aparc.annot.new



Pingback: Displaying vertexwise and facewise brain maps | Brainder.
Pingback: Splitting the cortical surface into independent regions | Brainder.
Thanks for this blog. You saved this neuroscientist a lot of time.
Hello, Thank you for this blog..
I followed the steps but I did not get the brain shown at the first of the article!
I get the following: ”
Found embedded color table in annotation
131716 vertices did not have an annotation!
surfer: single buffered window
surfer:tkoInitWindow(bert)
OpenGL Warning: Failed to connect to host. Make sure 3D acceleration is enabled for this VM”
Hi Ahmad,
The second message has to do with OpenGL not enable in your machine. Are you using this over VNC? Or a virtual machine? It’s something related to the local configuration it seems.
All the best,
Anderson
Hi! Did not know about the annotation files in Freesurfer. Thanks!
In case you want to convert from the labels descriptor of ITK-snap to the one of Freesurfer (and vice versa), you can check my tool to manipulate labels and descriptors:
https://github.com/SebastianoF/LABelsToolkit
The method is under the class LabelsDescriptorManager under the module LABelsToolkit/tools/descriptions/manipulate_descriptors.py
Thanks for sharing!
Thanks! I am devolping a web application that compare patologican patients with a normal database patients and with your post I could show the results using this approach .
Dear Anderson, What about mixing positive and negative values?
When I change a value in your example ods file to negative (in lh.data tab, e.g. -10 for parsorbitalis) and change the first line of the colormap tab to -1 -11.000 160 160 160 (gray) I get 160 160 160 for parsorbitalis in lh.aparc.annot.ctab tab.
Hi Joost,
The spreadsheet can be modified so as to make it more general. In particular, the formulas in column B of ‘colormap’ can be made more general and accommodate negative values — I suppose I should do that…
However, a quick workaround is to add a constant to your data that is simply abs(min(data)). Then it should work as is.
All the best,
Anderson